2020-10-09
10:15 - 11:15
Visual Computing Forum
Vidar R. Jensen is a Professor of Chemistry at the University of Bergen (http://jensen.uib.no/). His main interest is design of catalysts and other functional molecules. In recent years he has developed and used in silico methods to design new catalysts that have subsequently found practical use. Prof. Jensen envisions a future in which molecular and materials discovery is largely realized via “Big Computing / Big Data”, not serendipity.
Abstract:
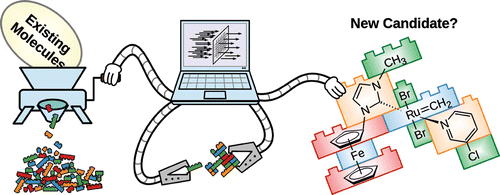
The contributions of the computational molecular sciences to improved health and sustainability are growing rapidly. This growth is spurred by spectacular advances in algorithms and computational infrastructure, which allow atom-scale modeling of complex chemical and biochemical systems. The calculations offer invaluable microscopic insight into how these systems work. However, molecular simulations may offer much more: They may, when combined with automated combinatorial or heuristic methods, speed up discovery of pharmaceuticals and other functional molecules and materials and reduce the reliance on costly wet-lab experiments. In other words, these compounds and materials are discovered largely thanks to powerful combinations of cheminformatics methods for high-throughput virtual screening and design with those of atomistic modeling. Here, we will describe a cheminformatics-based method that automatically assembles, tests, and designs molecules from
building blocks following user-defined criteria. The method stands out by being the most general and widely applicable of the so-called de novo design methods. A Java-based computer code is freely available (https://github.com/denoptim-project/DENOPTIM).